A recent study provides a new approach to identify aging brain cells in Alzheimer’s Disease.

Reference: Dehkordi SK, Jamie Walker, Eric Sah, Emma Bennett, Farzaneh Atrian, Bess Frost, Benjamin Woost, Rachel E. Bennett, Timothy C. Orr, Yingyue Zhou, Prabhakar S. Andhey, Marco Colonna, Peter H. Sudmant, Peng Xu, Minghui Wang, et al. Profiling senescent cells in human brains reveals neurons with CDKN2D/p19 and tau neuropathology. Nature Aging 2021; 1:1107-16.
Our bodies are not meant to last forever: becoming older and deteriorating with age is inevitable. Advanced age is accompanied by an increased risk for numerous diseases, including Alzheimer’s disease (AD). AD is a neurodegenerative disease, a type of disease in which the cells of the brain or spinal cord are progressively damaged and die. Because of the loss of brain cells, AD is primarily known for memory loss and decline in other mental capabilities. However, it is still unclear what causes AD. What is happening to our cells as we age? How do cellular changes contribute to AD and other age-related conditions? Can we target these changes to turn back the clock, or at least slow it down?
What is driving aging and age-related diseases?
When a cell stops functioning properly, it is programmed to die, and the remnants are digested by the immune system. However, cells will sometimes enter a state called cellular senescence in which the cell stops responding to signals but remains in the body. Senescent cells are characterized by DNA damage and the release of inflammatory factors that can damage the surrounding tissue1. As we age, our bodies accumulate more and more senescent cells2, 3. However, cellular senescence is emerging as a culprit in many conditions associated with aging, including neurodegenerative, cardiovascular, and metabolic diseases4-6. On a positive note, this means that there is potential to target senescence as a treatment for these diseases5, 7, 8, but identifying senescent cells is challenging. A new study by Dehkordi and colleagues provides a new approach for screening and identifying senescent brain cells in patients with AD9.
Using a new bioinformatics approach to define senescent cells: An ongoing challenge
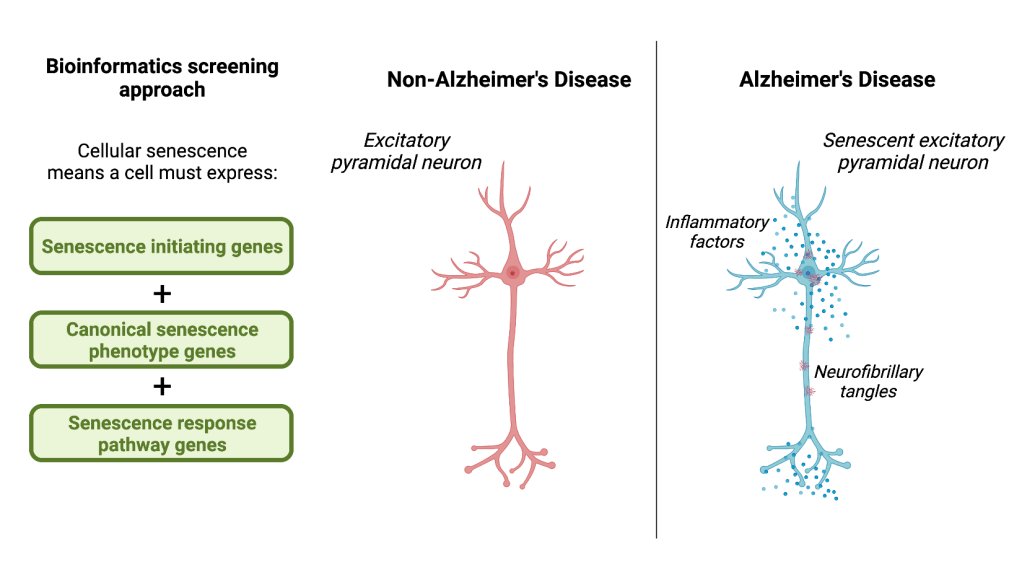
Currently, identifying senescent cells is complicated because there are inconsistencies in how these cells are defined1, 10, 11. To address this problem, Dehkordi and colleagues developed three bioinformatics tools called eigengenes to screen for senescent cells. Each eigengene contains a set of multiple genes known to be involved in a different stage of cellular senescence:
- Senescence initiating – Genes that are activated when the cell is undergoing stress but not yet in a senescent cell state.
- Canonical senescence phenotype – Genes that are increased to stop the cell from responding to signals and continuing with cell growth. These genes are widely accepted as markers of senescence.
- Senescence response pathway – Genes related to the release of inflammatory factors that occurs in parallel with the senescent cell state.
Within each eigengene, the authors looked for changes in the expression of these characteristic genes in different cell types in a brain region called the dorsolateral prefrontal cortex; a region crucial for numerous mental processes, such as working memory, decision-making, self-regulation, etc. Importantly, cells needed to meet each criterion within all three eigengenes to be ruled as “senescent.”
Senescence contributes to features of Alzheimer’s Disease
Upon examining cells from the dorsolateral prefrontal cortex, Dehkordi and colleagues found that cellular senescence in AD patients’ brains predominantly affects excitatory pyramidal neurons. Excitatory pyramidal neurons are brain cells that regulate communication with other parts of the brain and body and are involved in cognitive functions. Senescence in excitatory pyramidal neurons means that these cells cannot perform their functions, and they may be harming neighboring cells by increasing inflammation in the surrounding area. After determining the cell types affected in the AD brain, Dehkordi and colleagues explored how excitatory pyramidal neurons became senescent. The authors found that expression of the gene CDKN2D, which produces a protein called p19, was highly increased in senescent neurons relative to non-senescent neurons. P19 protein regulates the expression of our genetic material for cell growth to progress, meaning that excess p19 makes too much of our genetic material inaccessible and prevents the cell from carrying out its functions. Furthermore, Dehkordi and colleagues demonstrated a potential connection between senescence and one of the hallmark features of AD, neurofibrillary tangles. Neurofibrillary tangles are the accumulation of misfolded proteins that occurs within neurons during AD. The authors found that cells with neurofibrillary tangles in patients with AD also expressed p19, and these p19/ neurofibrillary tangle-containing cells displayed an irregular cell shape. In other words, senescent cells express neurofibrillary tangles, and neurofibrillary tangle-containing neurons become senescent.
What does it all mean?
All in all, the work by Dehkordi and colleagues provides important insights into potential mechanisms of AD based on the interactions that were observed between p19 and neurofibrillary tangles, although future exploration is required. Importantly, the work by Dehkordi and colleagues provides a better characterization of senescent cells by using a novel approach for defining and screening senescent cells. This approach holds implications for using senescence for therapeutic purposes applicable not only to AD but to other age-related conditions. The authors’ approach to using a variety of markers expressed during different levels of senescence advances our understanding of the senescent cell state. However, because of their study design, the conclusions of the paper may be biased. The authors may be overlooking cells with unique senescent profiles that do not fit within the specific parameters that they chose. Nonetheless, the findings by Dehkordi and colleagues provide an important step towards understanding brain cellular senescence in age-related diseases.
Additional References
1. Gorgoulis V, P. D. Adams, A. Alimonti, D. C. Bennett, O. Bischof, C. Bishop, J. Campisi, M. Collado, K. Evangelou, G. Ferbeyre, J. Gil, E. Hara, V. Krizhanovsky, D. Jurk, A. B. Maier, et al. Cellular senescence: Defining a path forward.Cell Elsevier Inc; 2019; 179:813-27.
2. Yousefzadeh MJ, J. Zhao, C. Bukata, E. A. Wade, S. J. McGowan, L. A. Angelini, M. P. Bank, A. U. Gurkar, C. A. McGuckian, M. F. Calubag, J. I. Kato, C. E. Burd, P. D. Robbins and L. J. Niedernhofer. Tissue specificity of senescent cell accumulation during physiologic and accelerated aging of mice. Aging Cell.The Authors. Aging Cell published by the Anatomical Society and John Wiley & Sons Ltd; 2020; 19:e13094.
3. Almanzar N, Jane Antony, Ankit S. Baghel, Isaac Bakerman, Ishita Bansal, Ben A. Barres, Philip A. Beachy, Daniela Berdnik, Biter Bilen, Douglas Brownfield, Corey Cain, Charles K. F. Chan, Michelle B. Chen, Michael F. Clarke, Stephanie D. Conley, et al. A single-cell transcriptomic atlas characterizes ageing tissues in the mouse. Nature 2020; 583:590-5. https://doi.org/10.1038/s41586-020-2496-1.
4. Kritsilis M, S. V Rizou, P. N. Koutsoudaki, K. Evangelou, V. G. Gorgoulis and D. Papadopoulos. Ageing, cellular senescence and neurodegenerative disease. Int.J.Mol.Sci. 2018; 19:2937. doi: 10.3390/ijms19102937.
5. Childs BG, H. Li and J. M. van Deursen. Senescent cells: A therapeutic target for cardiovascular disease. J.Clin.Invest. 2018; 128:1217-28.
6. Sikora E, Anna Bielak-Zmijewska and Grazyna Mosieniak. Cellular senescence in ageing, age-related disease and longevity. Current Vascular Pharmacology 2013; 12:698-706.
7. Zhu Y, T. Tchkonia, T. Pirtskhalava, A. C. Gower, H. Ding, N. Giorgadze, A. K. Palmer, Y. Ikeno, G. B. Hubbard, M. Lenburg, S. P. O’Hara, N. F. LaRusso, J. D. Miller, C. M. Roos, G. C. Verzosa, et al. The achilles’ heel of senescent cells: From transcriptome to senolytic drugs. Aging Cell.The Authors. Aging Cell published by the Anatomical Society and John Wiley & Sons Ltd; 2015; 14:644-58.
8. Baker DJ, T. Wijshake, T. Tchkonia, N. K. LeBrasseur, B. G. Childs, B. van de Sluis, J. L. Kirkland and Deursen, J. M. A. van. Clearance of p16Ink4a-positive senescent cells delays ageing-associated disorders. Nature 2011; 479:232-6. https://www.narcis.nl/publication/RecordID/oai:repository.ubn.ru.nl:2066%2F96729.
9. Dehkordi SK, Jamie Walker, Eric Sah, Emma Bennett, Farzaneh Atrian, Bess Frost, Benjamin Woost, Rachel E. Bennett, Timothy C. Orr, Yingyue Zhou, Prabhakar S. Andhey, Marco Colonna, Peter H. Sudmant, Peng Xu, Minghui Wang, et al. Profiling senescent cells in human brains reveals neurons with CDKN2D/p19 and tau neuropathology. Nature Aging 2021; 1:1107-16. https://doi.org/10.1038/s43587-021-00142-3.
10. Hayflick L and P. S. Moorhead. The serial cultivation of human diploid cell strains.Exp.Cell Res. 1961; 25:585-621.
11. Jurk D, Chunfang Wang, Satomi Miwa, Mandy Maddick, Viktor Korolchuk, Avgi Tsolou, Efstathios S. Gonos, Christopher Thrasivoulou, M. Jill Saffrey, Kerry Cameron and Thomas von Zglinicki. Postmitotic neurons develop a p21‐dependent senescence‐like phenotype driven by a DNA damage response. Aging cell Blackwell Publishing Ltd; 2012; 11:996-1004. https://onlinelibrary.wiley.com/doi/abs/10.1111/j.1474-9726.2012.00870.x.
