A recent study finds that improper protein formation triggers the accumulation of misfolded proteins and cellular stress in the brain cells of autism-like mice.

Created with Canva.com

Reference: Taketomi, T., Yasuda, T., Morita, R. et al. (2022). Autism-associated mutation in Hevin/Sparcl1 induces endoplasmic reticulum stress through structural instability. Scientific Reports, 12, 11891.
Over the last two decades, the number of children diagnosed with autism, a complex developmental disability affecting behavioral, social, and communication skills, has been steadily increasing2. Although there are many likely causes of this increased prevalence, extensive research has suggested that there is a large genetic component to autism3, 4, 5. Hundreds of genes have been identified as potential contributors in autism6, 7, 8. However, the specific genes that contribute to autism varies in each person. This variability makes it challenging to identify common pathways and provide effective therapeutic options. In other words, autism can be driven by either a mutation in a single gene or a unique combination of mutations in multiple genes9.
One such gene is Usp15, a gene responsible for making a protein (USP15) that regulates the formation of many other proteins. Under typical conditions, the instructions for making proteins are read and executed in a cell structure called the endoplasmic reticulum (ER). Newly made proteins are then transported to their target locations in the cell to carry out their jobs. However, chronic stress to the cell causes the ER to make proteins with structural issues – termed unfolded or misfolded proteins. These improperly made proteins remain in the ER, causing the ER to become overwhelmed and triggering what is known as ER stress. Importantly, deficiency in USP15 has been shown to be associated with ER stress10. Although mutations in Usp15 have also been linked to autism11, 12, whether USP15 deficiency leads to ER stress in autism remains unknown. Thus, Taketomi and colleagues addressed this gap by exploring mechanisms linking ER stress to USP15 deficiency, an interaction that may impact brain development in autism.
Autism-like brain cells contain mutant Hevin protein, causing ER stress
People with autism commonly experience changes in the processes that control brain development, leading to alterations in their brain structure13. An autism-associated protein, Hevin, is important for several essential processes of healthy brain development. Therefore, Taketomi and colleagues hypothesized that Hevin protein may be affected by USP15-deficiency in autism.
To examine how USP15 deficiency leads to instability in the structure of Hevin protein, the authors isolated brain cells from a mouse model of autism that lacks USP15. From these cells, they first isolated the instructions for Hevin protein formation and found that a portion of these instructions was either missing or fused with another sequence of instructions. Using structural analyses, they found that the final structure of Hevin protein cannot fold properly because of these faulty instructions. Furthermore, because the protein is lacking a crucial element, Hevin cannot relocate to where it is supposed to be in the cell. Instead, misfolded Hevin protein builds up in the ER. By measuring the levels of stress response markers, the authors found that this accumulation of mutant Hevin protein triggers the ER stress response in USP15-deficient brain cells. Therefore, a lack of USP15 leads to misfolded Hevin protein and ER stress in brain cells.
Similar Hevin mutation in humans with autism also triggers ER stress
The findings of the authors prompt the question: Do autistic humans have similar genetic mutations as the autism-like mice and experience the same ER stress mechanism in their brain cells? Indeed, there is a specific mutation in humans with autism that also affects the Hevin protein. To discover whether this human Hevin protein variant triggers the same ER stress mechanism, the authors created another model containing the human mutation in Hevin. Similar to their initial results, they found that the human version of mutant Hevin protein cannot fold properly and triggers the same ER stress response as the USP15-deficient mice.
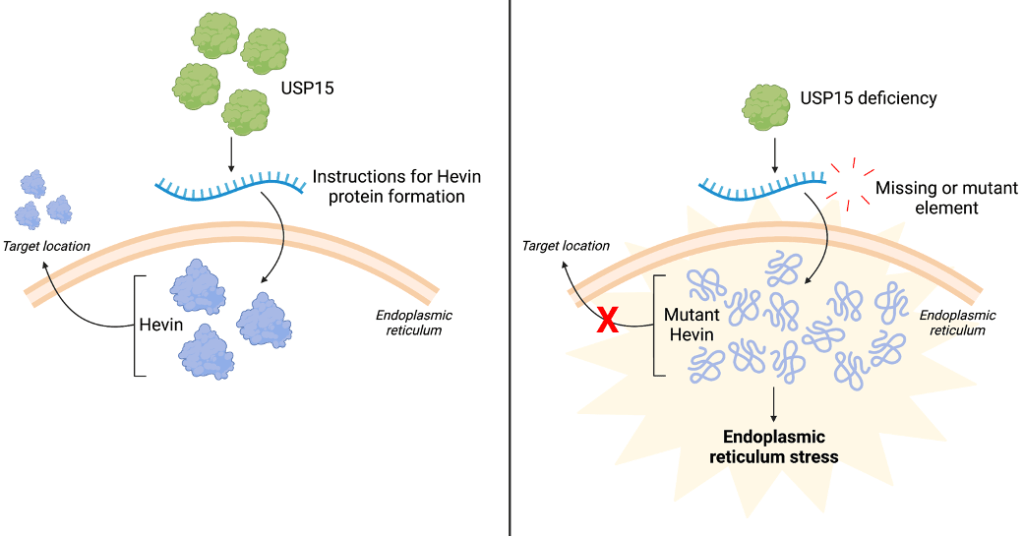
Created with BioRender.com
(On the left) Under typical circumstances, USP15 protein (green) regulates the instructions (blue) to make Hevin protein (purple), which is made in the endoplasmic reticulum. Once made, Hevin protein uses the location information in its instructions to relocate to complete its assigned job. However, USP15 deficiency (on the right), which can occur during autism, can lead to misregulation of Hevin protein. Instructions that are missing or have a mutated component are used to make Hevin protein. This leads to misfolded Hevin protein (purple lines) that becomes trapped in the endoplasmic reticulum, triggering endoplasmic reticulum stress.
What does it all mean?
Taketomi et al. have identified a novel mechanism underlying autism: USP15 deficiency leads to structural issues in Hevin protein and subsequent ER stress in brain cells during autism. Considering the importance of Hevin in brain cell maturation, impaired Hevin protein may impact brain development and cell functioning in autistic people.
Autism presentation is heterogeneous, meaning that people who are seeking therapeutic options may require care that is tailored to their specific needs. However, studying all rare mutations in autism and identifying common pathways is challenging. An important consideration of this study is that the specific Hevin variants caused by USP15 deficiency in the current study may only be applicable to a small subset of autism cases. Given that USP15 regulates the production of many different proteins, further exploration is needed to identify additional proteins affected by USP15 deficiency. Changes in USP15 function will affect many proteins involved in various cellular and physiological processes, which could explain some of the diversity of autism presentation. Nonetheless, the findings of this new study provide an important pathway for an autism-associated mutation and paves the way for future studies to explore how ER stress in brain cells changes brain development during autism.
Additional References:
- Taketomi, T., Yasuda, T., Morita, R., Kim, J., Shigeta, Y., Eroglu, C., Harada, R., & Tsuruta, F. (2022). Autism-associated mutation in Hevin/Sparcl1 induces endoplasmic reticulum stress through structural instability. Scientific reports, 12(1), 11891. https://doi.org/10.1038/s41598-022-15784-5
- Data & Statistics on Autism Spectrum Disorder. (2022, March 2). Centers for Disease Control and Prevention. https://www.cdc.gov/ncbddd/autism/data.html
- Ronald, A., Happé, F., Price, T. S., Baron-Cohen, S., & Plomin, R. (2006). Phenotypic and genetic overlap between autistic traits at the extremes of the general population. Journal of the American Academy of Child and Adolescent Psychiatry, 45(10), 1206–1214. https://doi.org/10.1097/01.chi.0000230165.54117.41
- Rylaarsdam, L., & Guemez-Gamboa, A. (2019). Genetic Causes and Modifiers of Autism Spectrum Disorder. Frontiers in cellular neuroscience, 13, 385. https://doi.org/10.3389/fncel.2019.00385
- Constantino, J. N., & Todd, R. D. (2003). Autistic traits in the general population: a twin study. Archives of general psychiatry, 60(5), 524–530. https://doi.org/10.1001/archpsyc.60.5.524
- Grove, J., Ripke, S., Als, T. D., Mattheisen, M., Walters, R. K., Won, H., Pallesen, J., Agerbo, E., Andreassen, O. A., Anney, R., Awashti, S., Belliveau, R., Bettella, F., Buxbaum, J. D., Bybjerg-Grauholm, J., Bækvad-Hansen, M., Cerrato, F., Chambert, K., Christensen, J. H., Churchhouse, C., … Børglum, A. D. (2019). Identification of common genetic risk variants for autism spectrum disorder. Nature genetics, 51(3), 431–444. https://doi.org/10.1038/s41588-019-0344-8
- Matoba, N., Liang, D., Sun, H., Aygün, N., McAfee, J. C., Davis, J. E., Raffield, L. M., Qian, H., Piven, J., Li, Y., Kosuri, S., Won, H., & Stein, J. L. (2020). Common genetic risk variants identified in the SPARK cohort support DDHD2 as a candidate risk gene for autism. Translational psychiatry, 10(1), 265. https://doi.org/10.1038/s41398-020-00953-9
- Fu, J. M., Satterstrom, F. K., Peng, M., Brand, H., Collins, R. L., Dong, S., Wamsley, B., Klei, L., Wang, L., Hao, S. P., Stevens, C. R., Cusick, C., Babadi, M., Banks, E., Collins, B., Dodge, S., Gabriel, S. B., Gauthier, L., Lee, S. K., Liang, L., … Talkowski, M. E. (2022). Rare coding variation provides insight into the genetic architecture and phenotypic context of autism. Nature genetics, 54(9), 1320–1331. https://doi.org/10.1038/s41588-022-01104-0
- O’Roak, B. J., Vives, L., Fu, W., Egertson, J. D., Stanaway, I. B., Phelps, I. G., Carvill, G., Kumar, A., Lee, C., Ankenman, K., Munson, J., Hiatt, J. B., Turner, E. H., Levy, R., O’Day, D. R., Krumm, N., Coe, B. P., Martin, B. K., Borenstein, E., Nickerson, D. A., … Shendure, J. (2012). Multiplex targeted sequencing identifies recurrently mutated genes in autism spectrum disorders. Science (New York, N.Y.), 338(6114), 1619–1622. https://doi.org/10.1126/science.1227764
- Kim, J., Nakamura, J., Hamada, C., Taketomi, T., Yano, S., Okajima, T., Kashiwabara, S. I., Baba, T., Sato, B., Chiba, T., & Tsuruta, F. (2020). USP15 Deubiquitinates TUT1 Associated with RNA Metabolism and Maintains Cerebellar Homeostasis. Molecular and cellular biology, 40(21), e00098-20. https://doi.org/10.1128/MCB.00098-20
- Chen, R., Davis, L. K., Guter, S., Wei, Q., Jacob, S., Potter, M. H., Cox, N. J., Cook, E. H., Sutcliffe, J. S., & Li, B. (2017). Leveraging blood serotonin as an endophenotype to identify de novo and rare variants involved in autism. Molecular autism, 8, 14. https://doi.org/10.1186/s13229-017-0130-3
- Krupp, D. R., Barnard, R. A., Duffourd, Y., Evans, S. A., Mulqueen, R. M., Bernier, R., Rivière, J. B., Fombonne, E., & O’Roak, B. J. (2017). Exonic Mosaic Mutations Contribute Risk for Autism Spectrum Disorder. American journal of human genetics, 101(3), 369–390. https://doi.org/10.1016/j.ajhg.2017.07.016
- Hashem, S., Nisar, S., Bhat, A. A., Yadav, S. K., Azeem, M. W., Bagga, P., … & Haris, M. (2020). Genetics of structural and functional brain changes in autism spectrum disorder. Translational Psychiatry, 10(1), 1-17.
